Our Reseach
Problem
The current COVID pandemic demonstrates how infectious diseases collectively represent tremendous and unpredictable challenges for humankind and how novel drugs are urgently needed to continually improve and replenish our arsenal of therapeutics. The rate at which lead molecules derived from high-throughput screens (HTS) are optimized for eventual human use (e.g. bench-to-bedside) is significantly hampered, as the modern drug discovery process is entirely reliant on the manual iteration of small molecule syntheses with biological assays. This dependence on skilled medicinal chemists makes medicinal chemistry incredibly time consuming and costly.
Research Philosophy
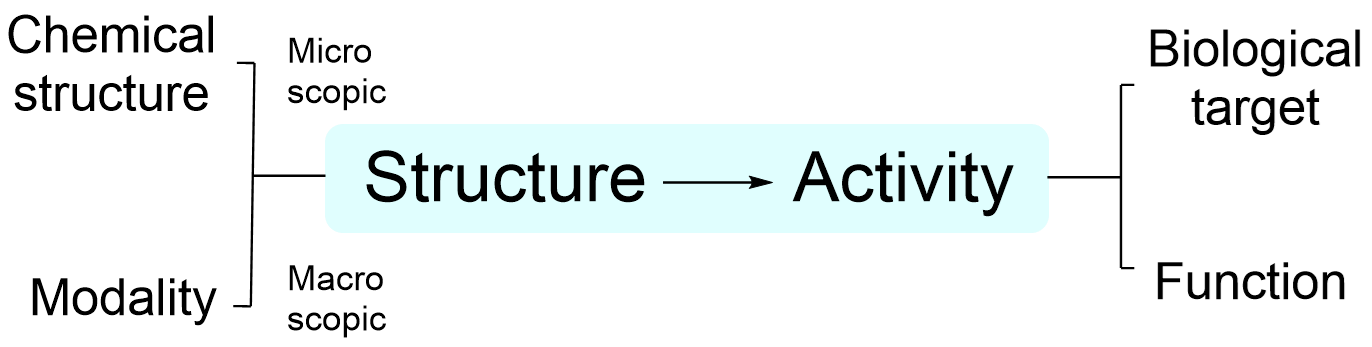
What is the central principle of medicinal chemistry? My answer is structure-activity relationship (SAR), while this paradigm must be further optimized to accelerate drug discovery. Our research program focusses on the development and expansion of each component of this SAR paradigm with an emphasis on targeting host-microbe interactions, with an ultimate goal to fundamentally change the medicinal chemistry and chemical biology.
Development of fundamental methodologies in medicinal chemistry
The increased accessibility of high-throughput screening methods and instruments has revolutionized how the scientific community searches for small molecule probes that modulate protein function and cellular pathways. Unfortunately, most initial screening hits require extensive optimization to yield molecules with the necessary potency and selectivity for biological applications. Lead optimization is a significant hurdle in the drug and probe development process.
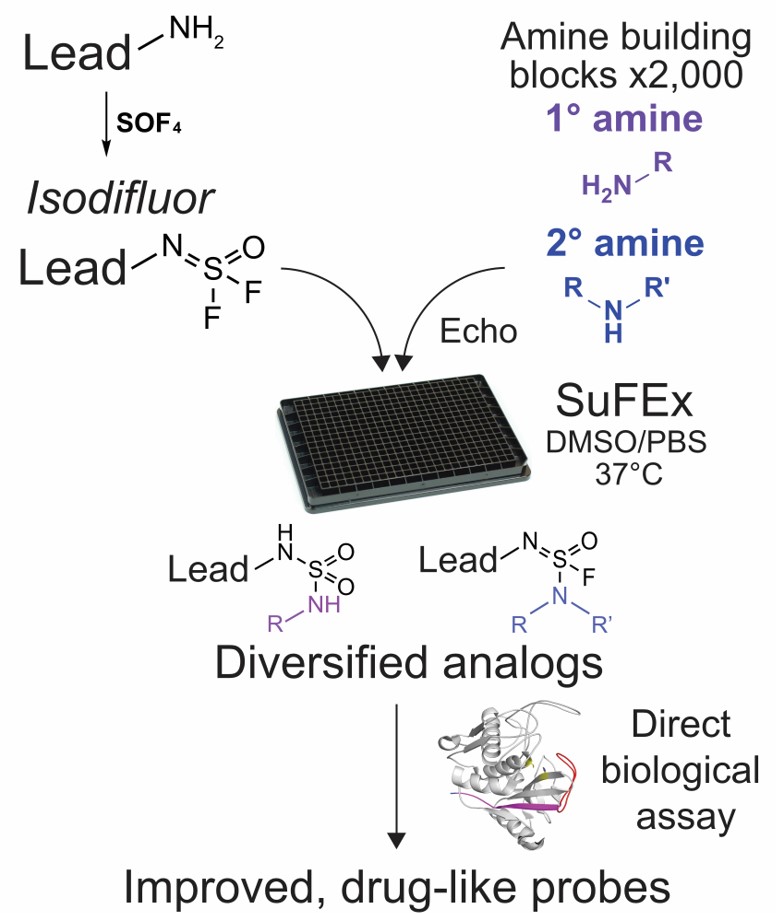
Our laboratory aims to develop and apply a universal HTMC methodology that could overcome these problems and accelerate the probe development. As a first step, we developed an SFx platform based on the recently developed click chemistry reaction sulfur(VI) fluoride exchange (SuFEx). The SFx platform uses a robotic liquid handler and ~2,000 amines to generate thousands of small molecule analogs overnight in picomole scale, followed by direct measurement of biological activity. The platform helped us generate improved, drug-like inhibitors of a virulence protease SpeB from S. pyogenes with >300-fold improved activity (IC50 = 29 nM) as well as inhibitors of an epigenetic regulator ENL with 200-fold improved cellular activity (EC50 = 63 nM). We are currently incorporating other chemical reactions, in particular click chemistry reactions into the platform to cover a wider chemical space in search of optimal ligands.
K99/R00: A universal high-throughput platform to improve biological functions of small molecules
Selected Publications
- SuFEx-Enabled High-Throughput Medicinal Chemistry. Kitamura et al., JACS. 2020; 10.1021/jacs.9b13652.
- Pharmacological inhibitors of the ENL YEATS domain. Erb, MA.; Wolan, DW.; Kitamura, S.; Chatterjee, A. Provisional patent. TSRI-014PR00
- Chemical inhibition of ENL/AF9 YEATS domains in acute leukemia. Garnar-Wortzel, L.*, Bishop, T.*, Kitamura, S.* (co-first author) et al., ACS Central Science. 2021; 10.1021/acscentsci.0c01550
- Antiviral small molecules that target the H1 hemagglutinin stem binding epitope. Wolan D.W., Wilson I.A., Yao Y., Kitamura S., Wu N.C., Kadam R.U., Lee C.-C.D.; TSRI-1361.276PRV.
Drug discovery and chemical biology of infectious diseases
We apply our high-throughput medicinal chemistry platform as well as traditional medicinal chemistry to pathogen targets to develop chemical probes useful for biological understanding of infectious diseases with direct therapeutic applications. Our targets include both infectious viruses and bacteria.
Selected Publications
- Rational design of potent and selective inhibitors of an epoxide hydrolase virulence factor from Pseudomonas aeruginosa. Kitamura et al. Journal of Medicinal Chemistry 2016. 10.1021/acs.jmedchem.6b00173
- Nanobody-based binding assay for the discovery of potent inhibitors of CFTR inhibitory factor (Cif). Vasylieva, N. et al. Analytica Chimica Acta. 2019;1057:106-113
- Lipoprotein signal peptidase inhibitors with antibiotic properties identified through design of a robust in vitro HT platform. Kitamura et al. Cell Chemical Biology 2017, 25, 301-308.e312.
- Diversity Oriented Clicking (DOC): Divergent synthesis of SuFExable pharmacophores from 2‐substituted‐alkynyl‐1‐sulfonyl fluoride (SASF) hubs. [MRSA active compounds] Smedley, C.J. et al., Angewandte Chemie International Edition 2020, doi: 10.1002/anie.202003219.
- An influenza A hemagglutinin small-molecule fusion inhibitor identified by a new high-throughput fluorescence polarization screen. Yao Y. et al. Proceedings of the National Academy of Sciences 2020, 202006893.
Chemical biology of host-microbiome interactions
Our body is colonized with a vast and diverse network of commensal bacteria that are essential for human metabolism, immune development and homeostasis, and protection from infiltration of pathogenic bacteria. We develop chemical probes and employ chemical biological techniques to rapidly and systematically identify host and microbial proteins associated with human diseases. The target proteins include both microbial proteins such as proteases as well as host protein targets important in the interaction with microbes.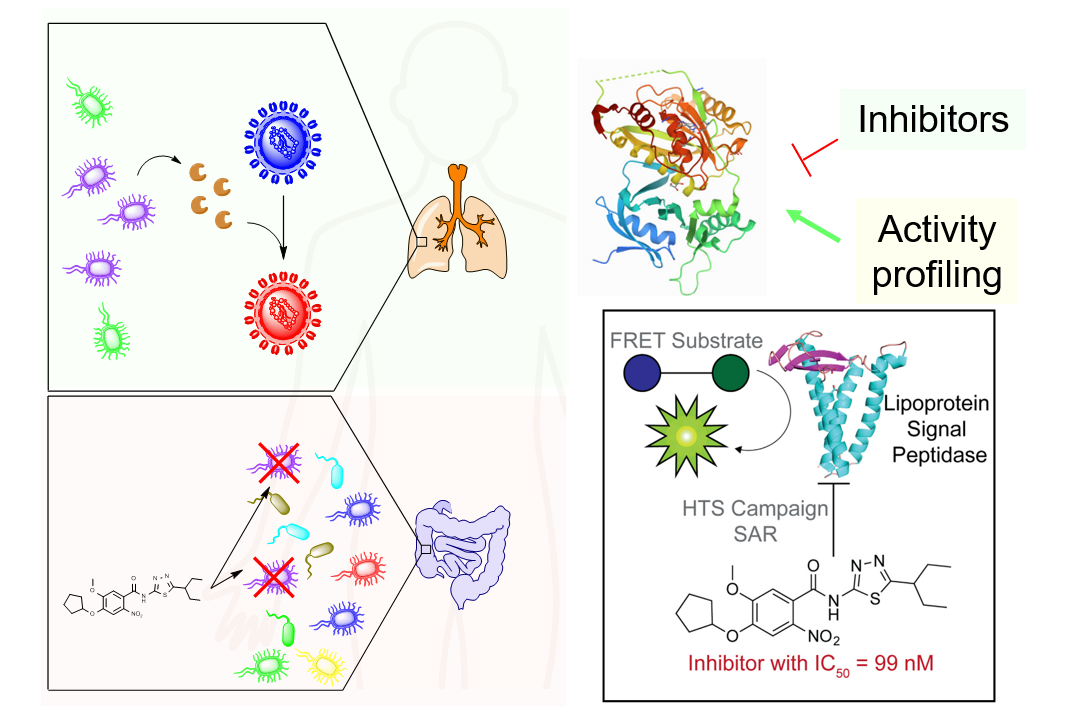
Selected Publications
- An irreversible inhibitor to probe the role of Streptococcus pyogenes cysteine protease SpeB in evasion of host complement defenses. Woehl, J.L.*; Kitamura, S.* (co-first author) et al. ACS Chemical Biology 2020, doi: 10.1021/acschembio.0c00191.
- Genetic basis for the cooperative bioactivation of plant lignans by Eggerthella lenta and other human gut bacteria. Bess, E. N. et al. Nature Microbiology 2020, 5, 56–66.
- A photo affinity probe that enriches folate binding proteins. Takamura, A. et al. Bioorganic & Medicinal Chemistry Letters 2020 10.1016/j.bmcl.2021.127903
- Metabolomics activity screening of T cell-induced colitis reveals novel anti-inflammatory metabolites. Montenegro-Burke, J.R. et al. Science Signaling 2021 10.1126/scisignal.abf6584
Expanding the modalities and function of molecules
Traditionally, inhibition of biological pathways was the major biological functions of small molecules while recently other functions are explored such as degradation or activation of target proteins. We aim to explore new modalities and functions of molecules. These efforts could enable us to modulate biological targets that are currently considered undruggable.
Selected publications
- Discovery of small-molecule enzyme activators by activity-based protein profiling. Kok et al. Nature Chemical Biology. 2020, doi: 10.1038/s41589-020-0555-4.
- Occurrence of urea-based soluble epoxide hydrolase inhibitors from the plants in the order Brassicales. Kitamura et al. PLoS One 2017, 12 (5), e0176571.
